Abstract
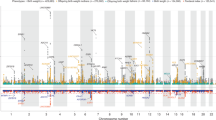
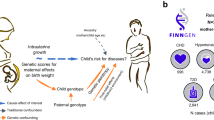
Birth weight variation is influenced by fetal and maternal genetic and non-genetic factors, and has been reproducibly associated with future cardio-metabolic health outcomes. In expanded genome-wide association analyses of own birth weight (n = 321,223) and offspring birth weight (n = 230,069 mothers), we identified 190 independent association signals (129 of which are novel). We used structural equation modeling to decompose the contributions of direct fetal and indirect maternal genetic effects, then applied Mendelian randomization to illuminate causal pathways. For example, both indirect maternal and direct fetal genetic effects drive the observational relationship between lower birth weight and higher later blood pressure: maternal blood pressure-raising alleles reduce offspring birth weight, but only direct fetal effects of these alleles, once inherited, increase later offspring blood pressure. Using maternal birth weight-lowering genotypes to proxy for an adverse intrauterine environment provided no evidence that it causally raises offspring blood pressure, indicating that the inverse birth weight–blood pressure association is attributable to genetic effects, and not to intrauterine programming.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The genotype and phenotype data are available on application from the UK Biobank (http://www.ukbiobank.ac.uk/). Individual cohorts participating in the EGG Consortium should be contacted directly as each cohort has different data access policies. GWAS summary statistics from this study are available via the EGG website (https://egg-consortium.org/).
Code availability
Custom-written code is available on request from N.M.W. (e-mail: n.warrington@uq.edu.au).
References
Barker, D. J. et al. Type 2 (non-insulin-dependent) diabetes mellitus, hypertension and hyperlipidaemia (syndrome X): relation to reduced fetal growth. Diabetologia 36, 62–67 (1993).
Martin-Gronert, M. S. & Ozanne, S. E. Mechanisms underlying the developmental origins of disease. Rev. Endocr. Metab. Disord. 13, 85–92 (2012).
Lumey, L. H., Stein, A. D. & Susser, E. Prenatal famine and adult health. Annu. Rev. Public Health 32, 237–262 (2011).
Ben-Shlomo, Y. & Smith, G. D. Deprivation in infancy or in adult life: which is more important for mortality risk? Lancet 337, 530–534 (1991).
Horikoshi, M. et al. Genome-wide associations for birth weight and correlations with adult disease. Nature 538, 248–252 (2016).
Hattersley, A. T. & Tooke, J. E. The fetal insulin hypothesis: an alternative explanation of the association of low birthweight with diabetes and vascular disease. Lancet 353, 1789–1792 (1999).
Beaumont, R. N. et al. Genome-wide association study of offspring birth weight in 86 577 women identifies five novel loci and highlights maternal genetic effects that are independent of fetal genetics. Hum. Mol. Genet. 27, 742–756 (2018).
Horikoshi, M. et al. New loci associated with birth weight identify genetic links between intrauterine growth and adult height and metabolism. Nat. Genet. 45, 76–82 (2013).
Freathy, R. M. et al. Variants in ADCY5 and near CCNL1 are associated with fetal growth and birth weight. Nat. Genet. 42, 430–435 (2010).
Hattersley, A. T. et al. Mutations in the glucokinase gene of the fetus result in reduced birth weight. Nat. Genet. 19, 268–270 (1998).
Eaves, L. J., Pourcain, B. S., Smith, G. D., York, T. P. & Evans, D. M. Resolving the effects of maternal and offspring genotype on dyadic outcomes in genome wide complex trait analysis (“M-GCTA”). Behav. Genet. 44, 445–455 (2014).
Warrington, N. M., Freathy, R. M., Neale, M. C. & Evans, D. M. Using structural equation modelling to jointly estimate maternal and fetal effects on birthweight in the UK Biobank. Int. J. Epidemiol. 47, 1229–1241 (2018).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 361–363 (2012).
GTEx Consortium. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Peng, S. et al. Expression quantitative trait loci (eQTLs) in human placentas suggest developmental origins of complex diseases. Hum. Mol. Genet. 26, 3432–3441 (2017).
Way, G. P., Youngstrom, D. W., Hankenson, K. D., Greene, C. S. & Grant, S. F. Implicating candidate genes at GWAS signals by leveraging topologically associating domains. Eur. J. Hum. Genet. 25, 1286–1289 (2017).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Zhang, G. et al. Genetic associations with gestational duration and spontaneous preterm birth. N. Engl. J. Med. 377, 1156–1167 (2017).
Smith, G. D. & Ebrahim, S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Smith, G. D. et al. Clustered environments and randomized genes: a fundamental distinction between conventional and genetic epidemiology. PLoS Med. 4, e352 (2007).
Tyrrell, J. et al. Genetic evidence for causal relationships between maternal obesity-related traits and birth weight. J. Am. Med. Assoc. 315, 1129–1140 (2016).
Pierce, B. L. & Burgess, S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. Am. J. Epidemiol. 178, 1177–1184 (2013).
Walton, A. & Hammond, J. The maternal effects on growth and conformation in shire horse–Shetland pony crosses. Proc. R. Soc. Lond. B 125, 311–335 (1938).
Smith, D. W. et al. Shifting linear growth during infancy: illustration of genetic factors in growth from fetal life through infancy. J. Pediatr. 89, 225–230 (1976).
Sorensen, T. et al. Comparison of associations of maternal peri-pregnancy and paternal anthropometrics with child anthropometrics from birth through age 7 y assessed in the Danish National Birth Cohort. Am. J. Clin. Nutr. 104, 389–396 (2016).
Hypponen, E., Power, C. & Smith, G. D. Parental growth at different life stages and offspring birthweight: an intergenerational cohort study. Paediatr. Perinat. Epidemiol. 18, 168–177 (2004).
Knight, B. et al. Evidence of genetic regulation of fetal longitudinal growth. Early Hum. Dev. 81, 823–831 (2005).
Nahum, G. G. & Stanislaw, H. Relationship of paternal factors to birth weight. J. Reprod. Med. 48, 963–968 (2003).
Griffiths, L. J., Dezateux, C. & Cole, T. J. Differential parental weight and height contributions to offspring birthweight and weight gain in infancy. Int. J. Epidemiol. 36, 104–107 (2007).
Wood, A. R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Zhang, G. et al. Assessing the causal relationship of maternal height on birth size and gestational age at birth: a Mendelian randomization analysis. PLoS Med. 12, e1001865 (2015).
Tyrrell, J. et al. Height, body mass index, and socioeconomic status: Mendelian randomisation study in UK Biobank. Br. Med. J. 352, i582 (2016).
Li, X., Redline, S., Zhang, X., Williams, S. & Zhu, X. Height associated variants demonstrate assortative mating in human populations. Sci. Rep. 7, 15689 (2017).
Pedersen J. Diabetes and Pregnancy: Blood Sugar of Newborn Infants. PhD thesis (Danish Science Press, 1952).
Metzger, B. E. et al. Hyperglycemia and adverse pregnancy outcomes. N. Eng. J. Med. 358, 1991–2002 (2008).
Crowther, C. A. et al. Effect of treatment of gestational diabetes mellitus on pregnancy outcomes. N. Eng. J. Med. 352, 2477–2486 (2005).
Jarvelin, M. R. et al. Early life factors and blood pressure at age 31 years in the 1966 northern Finland birth cohort. Hypertension 44, 838–846 (2004).
Tu, Y. K., West, R., Ellison, G. T. & Gilthorpe, M. S. Why evidence for the fetal origins of adult disease might be a statistical artifact: the “reversal paradox” for the relation between birth weight and blood pressure in later life. Am. J. Epidemiol. 161, 27–32 (2005).
Huxley, R., Neil, A. & Collins, R. Unravelling the fetal origins hypothesis: is there really an inverse association between birthweight and subsequent blood pressure? Lancet 360, 659–665 (2002).
Wang, T. et al. Low birthweight and risk of type 2 diabetes: a Mendelian randomisation study. Diabetologia 59, 1920–1927 (2016).
Freathy, R. M. Can genetic evidence help us to understand the fetal origins of type 2 diabetes? Diabetologia 59, 1850–1854 (2016).
Zanetti, D. et al. Birthweight, type 2 diabetes mellitus, and cardiovascular disease: addressing the Barker hypothesis with Mendelian randomization. Circ. Genom. Precis. Med. 11, e002054 (2018).
Lawlor, D. et al. Using Mendelian randomization to determine causal effects of maternal pregnancy (intrauterine) exposures on offspring outcomes: sources of bias and methods for assessing them. Wellcome Open Res. 2, 11 (2017).
Magi, R. & Morris, A. P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Kemp, J. P. et al. Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat. Genet. 49, 1468–1475 (2017).
Jones, S. E. et al. Genome-wide association analyses of chronotype in 697,828 individuals provides insights into circadian rhythms. Nat. Commun. 10, 343 (2019).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Ihaka, R. & Gentleman, R. R: a language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Mardia, K. V., Kent, J. T. & Bibby, J. M. Multivariate Analysis (Academic Press, 1979).
Segrè, A. V. et al. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Frayling, T. M. et al. A common allele in FGF21 associated with sugar intake is associated with body shape, lower total body-fat percentage, and higher blood pressure. Cell Rep. 23, 327–336 (2018).
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2016).
Prokopenko, I. et al. A central role for GRB10 in regulation of islet function in man. PLoS Genet. 10, e1004235 (2014).
Burgess, S., Scott, R. A., Timpson, N. J., Davey Smith, G. & Thompson, S. G. Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur. J. Epidemiol. 30, 543–552 (2015).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Delaneau, O., Zagury, J. F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Acknowledgements
Full acknowledgements and supporting grant details can be found in the Supplementary Information.
Author information
Authors and Affiliations
Consortia
Contributions
The central analysis and writing team comprised N.M.W., R.N.B., M.H., F.R.D., K.K.O., M.I.M., J.R.B.P., D.M.E. and R.M.F. Statistical analysis was performed by N.M.W., R.N.B., M.H., F.R.D., Ø.H., C.Lau., J.B., S.P., K.H., B.F., A.R.W., A.Mah., J.T., N.R.R., N.W.R., Z.Q., G-H.M., M.Vau., M.N., T.M.S., M.H.Z., J.P.B., N.G., M.N.K., R.L.-G., F.G., T.S.A., L.P., R.R., V.H., J.-J.H., L.-P.L., A.C., S.M., D.L.C., Y.W., E.T., C.A.W., C.T.H., N.V.-T., P.K.J., J.N.P., I.N., R.M., N.P., E.M.v.L., R.J., V.L., R.C.R., A.E., S.J.B., W.A., J.A.M., K.L.L., C.A., G.Z., L.J.M., J.Heik., A.H.C.v.K., B.D.C.v.S., K.J.G., N.R.v.Z., C.M.-G., Z.K., S.D., H.M., E.V.R.A., M.Mur., S.B.-G., D.M.H., J.M.Mer., K.E.S., P.A.L., S.E.M., B.M.S., J.-F.C., K.Pan., F.S., D.T., I.P., M.A.T., H.Y., K.S.R., S.E.J., P.-R.L., A.Mur., M.N.W., E.Z., G.V.D., Y.-Y.T., M.G.H., K.L.M., J.F.F., D.M.S., N.J.T., A.P.M., D.A.L., J.R.B.P., D.M.E. and R.M.F. Genotyping was performed by F.R.D., Ø.H., T.M.S., M.H.Z., N.G., R.L.-G., L.P., J.-J.H., L.-P.L., J.W.H., X.E., L.M., L.B., C.S.M., C.Lan., J.L., R.A.S., J.H.Z., G.H., S.M.R., A.J.B., J.F.-T., C.M.-G., H.G.d.H., F.R.R., Z.K., P.M.-V., H.M., E.V.R.A., M.Bus., M.A., P.K., M.Stu., T.A.L., C.M.v.D., A.K., E.Z., S.-M.S., G.W.M., H.C., J.F.W., T.G.M.V., C.E.P., E.E.W., T.D.S., T.L., P.V., H.B., K.B., J.C.M., F.R., J.F.F., T.H., O.P., A.G.U., M.-R.J., W.L.L., G.D.S., N.J.T., N.J.W., H.H., S.F.A.G., T.M.F., D.A.L., P.R.N., K.K.O., M.I.M., J.R.B.P., D.M.E. and R.M.F. Sample collection and phenotyping were performed by F.R.D., B.F., C.J.M., J.C., J.P.B., M.N.K., R.L.-G., F.G., R.R., I.N., H.M.I., J.W.H., L.S.-M., C.R., B.H., C.L.R., M.Kog., L.C., M.-F.H., C.S.M., F.D.M., C.Lan, J.L., R.A.S., J.H.Z., S.M.R., C.M.-G., H.G.d.H., Z.K., P.M.-V., S.D., G.W., M.M.-N., M.Sta., C.E.F., C.T., C.E.M.v.B., M.Bus., D.M.H., A.L., B.A.K., M.Bar., J.S., R.K.V., S.M.W., B.L.C., A.T., K.F.M., A.-M.E., T.A.L., A.K., H.N., K.Pah., O.T.R., B.J., G.V.D., S.-M.S., G.W.M., J.F.W., T.G.M.V., M.Vri., J.-C.H., L.J.B., C.E.P., L.S.A., J.B.B., J.G.E., E.E.W., A.T.H., T.D.S., M.Käh., J.S.V., T.L., P.V., H.B., K.B., M.Mel., E.A.N., D.O.M.-K., J.F.F., V.W.V.J., C.Pis., A.A.V., M.-R.J., C.Pow., E.H., W.L.L., G.D.S., N.J.W., H.H., S.F.A.G., D.A.L., K.K.O., M.I.M. and J.R.B.P. The study designers and principal investigators included J.P.B., I.N., H.M.I., L.S.-M., X.E., B.H., J.M.Mur., M.Kog., L.C., M.-F.H., F.D.M., M.A., A.T., M.Stu., K.F.M., A.-M.E., T.A.L., C.M.v.D., W.K., A.K., H.N., K.Pah., O.T.R., B.J., E.Z., G.V.D., Y.-Y.T., S.-M.S., G.W.M., H.C., J.F.W., T.G.M.V., M.Vri., E.J.C.N.d.G., H.N.K., J.-C.H., L.J.B., C.E.P., J.Hein., L.S.A., J.B.B., K.L.M., J.G.E., E.E.W., A.T.H., T.D.S., M.Käh., J.S.V., T.L., D.I.B., S.S., P.V., T.I.A.S., H.B., K.B., J.C.M., M.Mel., E.A.N., D.O.M.-K., F.R., A.H., J.F.F., V.W.V.J., T.H., C.Pis., A.A.V., O.P., A.G.U., M.-R.J., C.Pow., E.H., W.L.L., N.J.T., A.P.M., N.J.W., H.H., S.F.A.G., T.M.F., D.A.L., P.R.N., S.J., K.K.O., M.I.M., J.R.B.P. and R.M.F.
Corresponding authors
Ethics declarations
Competing interests
A.A.V. is an employee of AstraZeneca. S.F.A.G. has received support from GlaxoSmithKline for research that is not related to the study presented in this paper. D.A.L. has received support from Medtronic and Roche Diagnostics for biomarker research that is not related to the study presented in this paper. M.I.M. serves on advisory panels for Pfizer, Novo Nordisk and Zoe Global, has received honoraria from Merck, Pfizer, Novo Nordisk and Eli Lilly, has stock options in Zoe Global, and has received research funding from AbbVie, AstraZeneca, Boehringer Ingelheim, Eli Lilly, Janssen, Merck, Novo Nordisk, Pfizer, Roche, Sanofi–Aventis, Servier and Takeda.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary text and figures
Supplementary Figs. 1–19, Supplementary Tables 1–4, 16 and 18, and Supplementary Note
Supplementary Tables
Supplementary Tables 5–15, 17 and 19
Rights and permissions
About this article
Cite this article
Warrington, N.M., Beaumont, R.N., Horikoshi, M. et al. Maternal and fetal genetic effects on birth weight and their relevance to cardio-metabolic risk factors. Nat Genet 51, 804–814 (2019). https://doi.org/10.1038/s41588-019-0403-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0403-1
This article is cited by
-
Complex interplay of neurodevelopmental disorders (NDDs), fractures, and osteoporosis: a mendelian randomization study
BMC Psychiatry (2024)
-
Maternal plasma cortisol’s effect on offspring birth weight: a Mendelian Randomisation study
BMC Pregnancy and Childbirth (2024)
-
Intrauterine growth and the tangential expansion of the human cerebral cortex in times of food scarcity and abundance
Nature Communications (2024)
-
Parental and child genetic burden of glycaemic dysregulation and early-life cognitive development: an Asian and European prospective cohort study
Translational Psychiatry (2024)
-
Disentangling the link between maternal influences on birth weight and disease risk in 36,211 genotyped mother–child pairs
Communications Biology (2024)