Abstract
In mammals, the physiological activation of the glucocorticoid receptor (GR) by glucocorticoids (GCs) promotes the maturation of cardiomyocytes during late gestation, but the effect on postnatal cardiac growth and regenerative plasticity is unclear. Here we demonstrate that the GC–GR axis restrains cardiomyocyte proliferation during postnatal development. Cardiomyocyte-specific GR ablation in conditional knockout (cKO) mice delayed the postnatal cardiomyocyte cell cycle exit, hypertrophic growth and cytoarchitectural maturation. GR-cKO hearts showed increased expression of genes involved in glucose catabolism and reduced expression of genes promoting fatty acid oxidation and mitochondrial respiration. Accordingly, oxygen consumption in GR-cKO cardiomyocytes was less dependent on fatty acid oxidation, and glycolysis inhibition reverted GR-cKO effects on cardiomyocyte proliferation. GR ablation or transient pharmacological inhibition after myocardial infarction in juvenile and/or adult mice facilitated cardiomyocyte survival, cell cycle re-entry and division, leading to cardiac muscle regeneration along with reduced scar formation. Thus, GR restrains heart regeneration and may represent a therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Bongiovanni, C. et al. Reawakening the intrinsic cardiac regenerative potential: molecular strategies to boost dedifferentiation and proliferation of endogenous cardiomyocytes. Front. Cardiovasc. Med. 8, 750604 (2021).
Tzahor, E. & Poss, K. D. Cardiac regeneration strategies: staying young at heart. Science 356, 1035–1039 (2017).
Sadek, H. & Olson, E. N. Toward the goal of human heart regeneration. Cell Stem Cell 26, 7–16 (2020).
Eschenhagen, T. et al. Cardiomyocyte regeneration: a consensus statement. Circulation 136, 680–686 (2017).
van Berlo, J. H. & Molkentin, J. D. An emerging consensus on cardiac regeneration. Nat. Med. 20, 1386–1393 (2014).
Porrello, E. R. et al. Transient regenerative potential of the neonatal mouse heart. Science 331, 1078–1080 (2011).
Drenckhahn, J.-D. et al. Compensatory growth of healthy cardiac cells in the presence of diseased cells restores tissue homeostasis during heart development. Dev. Cell 15, 521–533 (2008).
Sampaio-Pinto, V. et al. Neonatal apex resection triggers cardiomyocyte proliferation, neovascularization and functional recovery despite local fibrosis. Stem Cell Rep. 10, 860–874 (2018).
Haubner, B. J. et al. Complete cardiac regeneration in a mouse model of myocardial infarction. Aging 4, 966–977 (2012).
Ye, L. et al. Early regenerative capacity in the porcine heart. Circulation 138, 2798–2808 (2018).
Zhu, W. et al. Regenerative potential of neonatal porcine hearts. Circulation 138, 2809–2816 (2018).
Li, Y. et al. Genetic tracing identifies early segregation of the cardiomyocyte and nonmyocyte lineages. Circ. Res. 125, 343–355 (2019).
Jopling, C. et al. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature 464, 606–609 (2010).
Kikuchi, K. et al. Primary contribution to zebrafish heart regeneration by gata4+ cardiomyocytes. Nature 464, 601–605 (2010).
Soonpaa, M. H., Kim, K. K., Pajak, L., Franklin, M. & Field, L. J. Cardiomyocyte DNA synthesis and binucleation during murine development. Am. J. Physiol. 271, H2183–H2189 (1996).
Li, F., Wang, X., Capasso, J. M. & Gerdes, A. M. Rapid transition of cardiac myocytes from hyperplasia to hypertrophy during postnatal development. J. Mol. Cell. Cardiol. 28, 1737–1746 (1996).
Bergmann, O. et al. Evidence for cardiomyocyte renewal in humans. Science 324, 98–102 (2009).
Senyo, S. E. et al. Mammalian heart renewal by pre-existing cardiomyocytes. Nature 493, 433–436 (2013).
Uygur, A. & Lee, R. T. Mechanisms of cardiac regeneration. Dev. Cell 36, 362–374 (2016).
Heallen, T. R., Kadow, Z. A., Kim, J. H., Wang, J. & Martin, J. F. Stimulating cardiogenesis as a treatment for heart failure. Circ. Res. 124, 1647–1657 (2019).
Hashimoto, H., Olson, E. N. & Bassel-Duby, R. Therapeutic approaches for cardiac regeneration and repair. Nat. Rev. Cardiol. 15, 585–600 (2018).
Cahill, T. J., Choudhury, R. P. & Riley, P. R. Heart regeneration and repair after myocardial infarction: translational opportunities for novel therapeutics. Nat. Rev. Drug Discov. 16, 699–717 (2017).
Galdos, F. X. et al. Cardiac regeneration: lessons from development. Circ. Res. 120, 941–959 (2017).
Oakley, R. H. & Cidlowski, J. A. Glucocorticoid signaling in the heart: a cardiomyocyte perspective. J. Steroid Biochem. Mol. Biol. 153, 27–34 (2015).
Richardson, R. V., Batchen, E. J., Denvir, M. A., Gray, G. A. & Chapman, K. E. Cardiac GR and MR: from development to pathology. Trends Endocrinol. Metab. 27, 35–43 (2016).
Rog-Zielinska, E. A., Richardson, R. V., Denvir, M. A. & Chapman, K. E. Glucocorticoids and foetal heart maturation; implications for prematurity and foetal programming. J. Mol. Endocrinol. 52, R125–R135 (2014).
Giraud, G. D., Louey, S., Jonker, S., Schultz, J. & Thornburg, K. L. Cortisol stimulates cell cycle activity in the cardiomyocyte of the sheep fetus. Endocrinology 147, 3643–3649 (2006).
Feng, X., Reini, S. A., Richards, E., Wood, C. E. & Keller-Wood, M. Cortisol stimulates proliferation and apoptosis in the late gestation fetal heart: differential effects of mineralocorticoid and glucocorticoid receptors. Am. J. Physiol. Regul. Integr. Comp. Physiol. 305, R343–R350 (2013).
de Vries, W. B. et al. Suppression of physiological cardiomyocyte proliferation in the rat pup after neonatal glucocorticosteroid treatment. Basic Res. Cardiol. 101, 36–42 (2006).
Gay, M. S., Li, Y., Xiong, F., Lin, T. & Zhang, L. Dexamethasone treatment of newborn rats decreases cardiomyocyte endowment in the developing heart through epigenetic modifications. PLoS ONE 10, e0125033 (2015).
Cutie, S., Payumo, A. Y., Lunn, D. & Huang, G. N. In vitro and in vivo roles of glucocorticoid and vitamin D receptors in the control of neonatal cardiomyocyte proliferative potential. J. Mol. Cell. Cardiol. 142, 126–134 (2020).
Tao, Z. et al. Dexamethasone inhibits regeneration and causes ventricular aneurysm in the neonatal porcine heart after myocardial infarction. J. Mol. Cell. Cardiol. 144, 15–23 (2020).
Hattori, F. et al. Nongenetic method for purifying stem cell-derived cardiomyocytes. Nat. Methods 7, 61–66 (2010).
Genangeli, M. et al. Development and application of a UHPLC–MS/MS method for the simultaneous determination of 17 steroidal hormones in equine serum. J. Mass Spectrom. 52, 22–29 (2017).
Morgan, R. A. et al. Dysregulation of cortisol metabolism in equine pituitary pars intermedia dysfunction. Endocrinology 159, 3791–3800 (2018).
Perogamvros, I., Ray, D. W. & Trainer, P. J. Regulation of cortisol bioavailability—effects on hormone measurement and action. Nat. Rev. Endocrinol. 8, 717–727 (2012).
Savu, L., Nunez, E. & Jayle, M. F. Corticosterone binding by mouse sera during foetal and post-natal development. Acta Endocrinol. 84, 177–190 (1977).
Oakley, R. H. et al. Essential role of stress hormone signaling in cardiomyocytes for the prevention of heart disease. Proc. Natl Acad. Sci. USA 110, 17035–17040 (2013).
Ali, H., Braga, L. & Giacca, M. Cardiac regeneration and remodelling of the cardiomyocyte cytoarchitecture. FEBS J. 287, 417–438 (2020).
Piquereau, J. & Ventura-Clapier, R. Maturation of cardiac energy metabolism during perinatal development. Front. Physiol. 9, 959 (2018).
Bae, J., Paltzer, W. G. & Mahmoud, A. I. The role of metabolism in heart failure and regeneration. Front. Cardiovasc. Med. 8, 702920 (2021).
Sim, C. B. et al. Sex-specific control of human heart maturation by the progesterone receptor. Circulation 143, 1614–1628 (2021).
Kubin, T. et al. Oncostatin M is a major mediator of cardiomyocyte dedifferentiation and remodeling. Cell Stem Cell 9, 420–432 (2011).
D’Uva, G. et al. ERBB2 triggers mammalian heart regeneration by promoting cardiomyocyte dedifferentiation and proliferation. Nat. Cell Biol. 17, 627–638 (2015).
Puente, B. N. et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 157, 565–579 (2014).
Honkoop, H. et al. Single-cell analysis uncovers that metabolic reprogramming by ErbB2 signaling is essential for cardiomyocyte proliferation in the regenerating heart. eLife 8, e50163 (2019).
Cardoso, A. C. et al. Mitochondrial substrate utilization regulates cardiomyocyte cell-cycle progression. Nat. Metab. 2, 167–178 (2020).
Mills, R. J. et al. Functional screening in human cardiac organoids reveals a metabolic mechanism for cardiomyocyte cell cycle arrest. Proc. Natl Acad. Sci. USA 114, E8372–E8381 (2017).
Cao, T. et al. Fatty acid oxidation promotes cardiomyocyte proliferation rate but does not change cardiomyocyte number in infant mice. Front. Cell Dev. Biol. 7, 42 (2019).
Severinova, E. et al. Glucocorticoid receptor-binding and transcriptome signature in cardiomyocytes. J. Am. Heart Assoc. 8, e011484 (2019).
Talman, V. et al. Molecular atlas of postnatal mouse heart development. J. Am. Heart Assoc. 7, e010378 (2018).
Parikh, S. S. et al. Thyroid and glucocorticoid hormones promote functional T-tubule development in human-induced pluripotent stem cell-derived cardiomyocytes. Circ. Res. 121, 1323–1330 (2017).
Karbassi, E. et al. Cardiomyocyte maturation: advances in knowledge and implications for regenerative medicine. Nat. Rev. Cardiol. 17, 341–359 (2020).
Hirose, K. et al. Evidence for hormonal control of heart regenerative capacity during endothermy acquisition. Science 364, 184–188 (2019).
Su, Q.-Q., Huang, X.-L., Qin, J. & Liu, Q.-S. Assessment of effects of mifepristone administration to lactating mice on the development and fertility of their progeny. J. Obstet. Gynaecol. Res. 41, 575–581 (2015).
Acknowledgements
This project was supported by the European Union’s Horizon 2020 Research and Innovation Program under the ERA-NET on Cardio Vascular Diseases (ERA-CVD) co-fund action to G.D. and E.T. (grant no. JCT2016-40-080); by Fondazione Luisa Fanti Melloni to G.D.; by Fondazione Carisbo to G.D. (grant no. 2020.0389); by the International Society for Heart Research to N.P. (ISHR research fellowship 2019); by University of Bologna AlmaIdea Junior Grant INTACT to L.I.; by University of Bologna AlmaIdea Junior Grant to M.L.; and by Ministry of Health - Ricerca Corrente - IRCCS MultiMedica. We thank Fondazione Del Monte (Bologna, Italy) and Centro Studi della Barbariga (Noventa Padovana, Padua, Italy) for the financial support finalized to A.M.P. for the acquisition of Seahorse XFe96 and JASCO V550 spectrophotometer instruments, respectively. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank G. Pelosi, D. Micello, E. Paiola, C. Kluc, A. Perlasca and B. Rainoldi for technical assistance in sectioning of paraffin-embedded samples. Part of this work was carried out in ALEMBIC, an advanced microscopy laboratory established by IRCCS Ospedale San Raffaele and Università Vita-Salute San Raffaele. We thank D. Zambroni for the technical assistance in the in vitro immunofluorescence imaging.
Author information
Authors and Affiliations
Contributions
N.P., F.S. and G.D. designed the experiments. N.P and F.S. carried out most of the experiments and analyzed the data. K.B.U., M.C. and G.D. performed myocardial infarction experiments and/or echocardiographic analyses. L.I. performed microrespirometry assays. S.D.P., C.B., C.M., F.P., E.P., R.S.P. and M.M. performed immunofluorescence, western blots and gene expression analysis. V.P. performed TEM analyses. L.B. performed the screening of FDA-approved GR agonists. R.T. helped with immunofluorescence image acquisition and time-lapse imaging. G.D. analyzed RNA sequencing data. G.C., A.M.P., M.L., C.V., M.G., R.R. and E.T. supervised the experiments done by their laboratory members, and G.D. supervised the entire project. N.P., F.S. and G.D. wrote the manuscript, with editing contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Richard Lee, Claude Libert and the other, anonymous, reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Circulating corticosterone levels in postnatal life and impact on cardiac muscle and stromal cell proliferation.
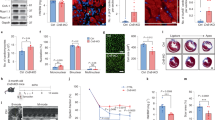
(a-b) Evaluation of 1-day-old (P1) cardiomyocyte proliferation by BrdU incorporation assay in (a) mixed cardiac cell cultures (n = 6 samples with a total of 2167 CMs analyzed) or (b) cardiomyocyte-enriched cultures (n = 6 samples with a total of 2650 CMs analyzed) following in vitro stimulation with corticosterone (CORT) at 10−8 M for 48 hours; CMs were identified by cTnI staining (see Methods for further details); (c) Quantification of DNA synthesis (BrdU) in postnatal day 1 (P1) cardiac stromal cells (n = 9 samples with a total of 4365 stromal cells analyzed) following administration of corticosterone (CORT) at 10−8 M or 10−5 M for 48 hours; (d) Evaluation of circulating corticosterone levels in the serum of postnatal day 1 (P1), postnatal day 3 (P3), postnatal day 5 (P5), postnatal day 7 (P7), postnatal day 14 (P14), postnatal day 28 (P28) and postnatal day 56 (P56) mice (n = 47 mice) by ELISA. In all panels, numerical data are presented as mean (error bars show s.e.m.); statistical significance was determined using two-sided Student’s t-test in a, b and one-way ANOVA followed by Tukey’s test in c, d.
Extended Data Fig. 2 Validation of separation between cardiomyocyte and stromal cell preparations.
(a-c) Cardiomyocytes (CMs) and stromal cells isolated from P1 and P7 hearts were analyzed by RT-PCR for markers of the major cardiac cell types (n = 12 mice), namely (a) cardiomyocytes (cTnT, cardiac troponin T), (b) endothelial cells (Pecam) and (c) fibroblasts (Ddr2), showing the absence of stromal cell markers in cardiomyocyte preparations. In all panels, numerical data are presented as mean (error bars show s.e.m.); statistical significance was determined using one-way ANOVA followed by Tukey’s test.
Extended Data Fig. 3 In vitro and in vivo analysis of cardiomyocyte-specific GR Knock Out (GR-cKO) mouse model.
(a) Immunofluorescence analysis of GR in cardiomyocyte cell cultures isolated from postnatal day 1 (P1) GR-cKO and control mice (403 CMs analyzed from n = 6 littermate mice sacrificed in a single session), showing the absence of GR expression in cardiomyocytes isolated from GR-cKO mice; arrows point at GR-positive and GR-negative nuclei in controls and GR-cKO cardiomyocytes, respectively; scale bars, 45 μm; (b) RT-PCR analysis of markers of cardiac pathological hypertrophy (Nppa, Nppb and Acta1) in postnatal day 7 (P7) GR-cKO and control heart lysates (n = 8 mice); (c) In vivo cardiomyocyte sarcomere status evaluation by immunofluorescence analysis of cardiac troponin T (cTnT) in P7 control (ctrl) and GR-cKO heart sections; images were obtained using a confocal microscope (n = 6 mice); scale bars, 15 μm for the main picture and 5 μm for the magnification; (d-g) Ultrastructure analysis by transmission electron microscopy (TEM) of P1 control (d,e) and GR-cKO (f,g) heart sections (n = 6 murine hearts isolated in two independent experiments), showing a slight more immature myofibrillar architecture in GR-cKO cardiomyocytes, characterized by less numerous myofibrils, sometimes randomly organized with cellular substructures (dotted lines in f compared to d), and isolated mitochondria (yellow arrows in f compared to d and magnification in g compared to e). Scale bars, 2 μm in (d, f) and 1 μm in (e, g). Legend for electron microscopy images: N = nucleus. (h) Heatmap of the top 5 statistically significant differentially expressed genes by RNA-Seq transcriptome analysis of P7 GR-cKO versus control hearts (n = 6 mice); the most statistically significant downregulated gene is Nr3c1 (GR); (i) Expression of de-differentiation markers (Runx1, Dab2 and Kit) in GR-cKO vs control P7 hearts from RNAseq analysis (n = 6 mice); (j) Cardiomyocyte dedifferentiation analysis by immunofluorescence staining for RUNX1/cTnT of P7 control (ctrl) and GR-cKO heart sections (n = 6 mice with a total of 17746 CMs analyzed); representative images are provided; scale bar, 20 μm; arrows point at dedifferentiated cardiomyocyte. In panel b and j, numerical data are presented as mean (error bars show s.e.m.); statistical significance was determined using two-sided Student’s t-test.
Extended Data Fig. 4 Heat maps of genes involved in ATP/lipids metabolism and mitochondria in postnatal day 7 GR-cKO versus controls hearts.
(a-b) Heat maps of statistically significant differentially expressed genes (pvalue < 0.05) by RNA-Seq transcriptome analysis of P7 GR-cKO versus control hearts (n = 6 mice) according to the following Biological Process gene ontology terms: (a) ATP metabolic process (GO:0046034) and (b) lipid metabolic process (GO:0006629); (c-d) Heat maps of statistically significant downregulated (c) or upregulated (d) genes (pvalue < 0.05) by RNA-Seq transcriptome analysis of P7 GR-cKO versus control hearts (n = 6 mice) according to Cellular Components gene ontology term ‘mitochondrion’ (GO:0005739).
Extended Data Fig. 5 Schematic diagram of energetic metabolic pathways modulated by GR in cardiomyocytes during early postnatal development.
Schematic diagram showing that GR reduces the expression of genes involved in glucose catabolism (via glycolysis and pentose phosphate pathway) while increasing the expression of genes involved in fatty acid oxidation and mitochondrial respiration.
Extended Data Fig. 6 Analysis of cellular respiration and activity of the respiratory complexes in GR-cKO versus control cardiomyocytes.
(a-c) Seahorse analysis of (a, n = 41 samples) basal respiration, (b, n = 6 samples) maximal respiration capacity and (c, n = 22 samples) extracellular acidification rate of cardiomyocyte cultures isolated from P1 control versus GR-cKO mice. (d) Specific activity of respiratory complexes (Complex I, CI; Complex II, CII; Complex III, CIII; Complex IV, CIV; Complex I+III, CI+III; Complex II+III, CII+III) normalized on citrate synthase (CS) activity, performed on mitochondria isolated from P7 GR-cKO and control hearts (n = 6 mice); (e-j) Seahorse analysis in response to (e-g) fatty acid oxidation inhibitor Etomoxir (Eto, 40 μM) or (h-j) inhibitor of glucose catabolism 2-deoxy-glucose (2dG, 5 μM) on P1 cardiomyocytes cultured in vitro (n = 7 samples in e, n = 8 samples in f, n = 8 samples in g; n = 7 samples in h; n = 7 samples in i; n = 7 samples in j); (k-m) Seahorse analysis of (k) acute response (n = 7 samples), (l) maximal respiration capacity (n = 7 samples) and (m) spare respiration capacity (n = 7 samples) in presence of 2-deoxy-glucose (2dG, 3 mM) in control versus GR-cKO P1 cardiomyocytes cultured in vitro. In all panels, numerical data are presented as mean (error bars show s.e.m.); statistical significance was determined using two-way ANOVA followed by Tukey’s test in h-j and two-sided Student’s t-test in l, m.
Extended Data Fig. 7 GR target genes upregulated during early postnatal development.
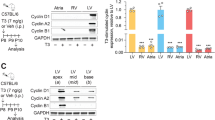
Heatmap of GR target genes significantly upregulated during early postnatal development. The list of genes whose expression is induced upon administration of dexamethasone, a synthetic GR agonist, by direct binding of GR to their promoter, was obtained from previous works (Severinova et al., 2019). The expression of these genes was evaluated in the cardiac tissue during early postnatal development, by employing publicly available data on postnatal day 1 (P1) to postnatal day 9 (P9) heart lysates (Talman et al., 2018). 15 out 51 genes were significantly upregulated from P1 to P9 (pvalue < 0.05).
Extended Data Fig. 8 Analysis of heart morphology and function in 4 months old GR-cKO versus control mice.
Echocardiographic measurements of ejection fraction (EF) of control and GR-cKO mice 15 weeks after birth (n = 14 mice); Numerical data are presented as mean (error bars show s.e.m.); statistical significance was determined using two-sided Student’s t-test.
Extended Data Fig. 9 GR transient inhibition facilitates cardiomyocyte proliferation and heart repair in the mouse model.
Schematic diagram showing the role of GR in controlling postnatal cardiomyocyte proliferation and heart regenerative ability. Cardiac GR abundance physiologically increases in cardiomyocytes during the first week of postnatal life (red line) and endogenous glucocorticoids/GR axis in cardiomyocytes contributes to the maturation of myofibrils-mitochondria organization coupled with a metabolic rewiring from glucose catabolism to fatty acid oxidation, in turn resulting in cardiomyocyte cell cycle exit and loss of cardiac regenerative ability. Transient inhibition of GR in adulthood facilitates cardiomyocyte proliferation and heart repair after damage.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2
Supplementary Video 1
Time-lapse movie of karyokinesis plus cytokinesis (cell division) in neonatal cardiomyocytes. This is a representative time-lapse video of neonatal cardiomyocytes performing karyokinesis plus cytokinesis (cell division). Heart cells were isolated from P1 mice, cultured in vitro for 48 hours and then labeled with TMRE (green) to identify cardiomyocytes (see Methods for further details) and imaged for 16 hours at 15-minute intervals.
Supplementary Video 2
Time-lapse movie of karyokinesis with no cytokinesis (bi-nucleation) in neonatal cardiomyocytes. This is a representative time-lapse video of P1 cardiomyocytes performing karyokinesis but not cytokinesis (bi-nucleation). Heart cells were isolated from P1 mice, cultured in vitro for 48 hours and then labeled with TMRE (green) to identify cardiomyocytes (see Methods for further details) and imaged for 16 hours at 15-minute intervals.
Supplementary Video 3
Time-lapse movie of karyokinesis plus cytokinesis (cell division) in GR-cKO neonatal cardiomyocytes. This is a representative time-lapse video of neonatal cardiomyocytes performing karyokinesis plus cytokinesis (cell division). Heart cells were isolated from P1 GR-cKO mice, cultured in vitro for 48 hours and then labeled with TMRE (green) to identify cardiomyocytes (see Methods for further details) and imaged for 16 hours at 15-minute intervals.
Supplementary Video 4
Time-lapse movie of karyokinesis with no cytokinesis (bi-nucleation) in GR-cKO neonatal cardiomyocytes. This is a representative time-lapse video of P1 cardiomyocytes performing karyokinesis but not cytokinesis (bi-nucleation). Heart cells were isolated from P1 GR-cKO mice, cultured in vitro for 48 hours and then labeled with TMRE (green) to identify cardiomyocytes (see Methods for further details) and imaged for 16 hours at 15-minute intervals.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 7
Statistical Source Data
Source Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 8
Statistical Source Data
Rights and permissions
About this article
Cite this article
Pianca, N., Sacchi, F., Umansky, K.B. et al. Glucocorticoid receptor antagonization propels endogenous cardiomyocyte proliferation and cardiac regeneration. Nat Cardiovasc Res 1, 617–633 (2022). https://doi.org/10.1038/s44161-022-00090-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-022-00090-0
This article is cited by
-
Thrombospondin 1 and Reelin act through Vldlr to regulate cardiac growth and repair
Basic Research in Cardiology (2023)
-
Glucocorticoid receptor inhibition promotes cardiac repair after MI
Nature Reviews Cardiology (2022)